Benchmarking Deep Learning and Vision Foundation Models for Atypical vs. Normal Mitosis Classification with Cross-Dataset Evaluation
Sweta Banerjee1 , Viktoria Weiss2
, Viktoria Weiss2 , Taryn A. Donovan3
, Taryn A. Donovan3 , Rutger H.J. Fick4, Thomas Conrad5, Jonas Ammeling6
, Rutger H.J. Fick4, Thomas Conrad5, Jonas Ammeling6 , Nils Porsche1, Robert Klopfleisch5
, Nils Porsche1, Robert Klopfleisch5 , Christopher C. Kaltenecker7
, Christopher C. Kaltenecker7 , Katharina Breininger8
, Katharina Breininger8 , Marc Aubreville1
, Marc Aubreville1 , Christof A. Bertram2
, Christof A. Bertram2
1: Flensburg University of Applied Sciences, Germany, 2: University of Veterinary Medicine, Vienna, Austria, 3: The Schwarzman Animal Medical Center, New York, USA, 4: Diffusely, Paris, France, 5: Freie Universität Berlin, Berlin, Germany, 6: Technische Hochschule Ingolstadt, Ingolstadt, Germany, 7: Medical University of Vienna, Austria, 8: Julius-Maximilians-Universität Würzburg, Würzburg, Germany
Publication date: 2026/03/12
https://doi.org/10.59275/j.melba.2026-6c1g
Abstract
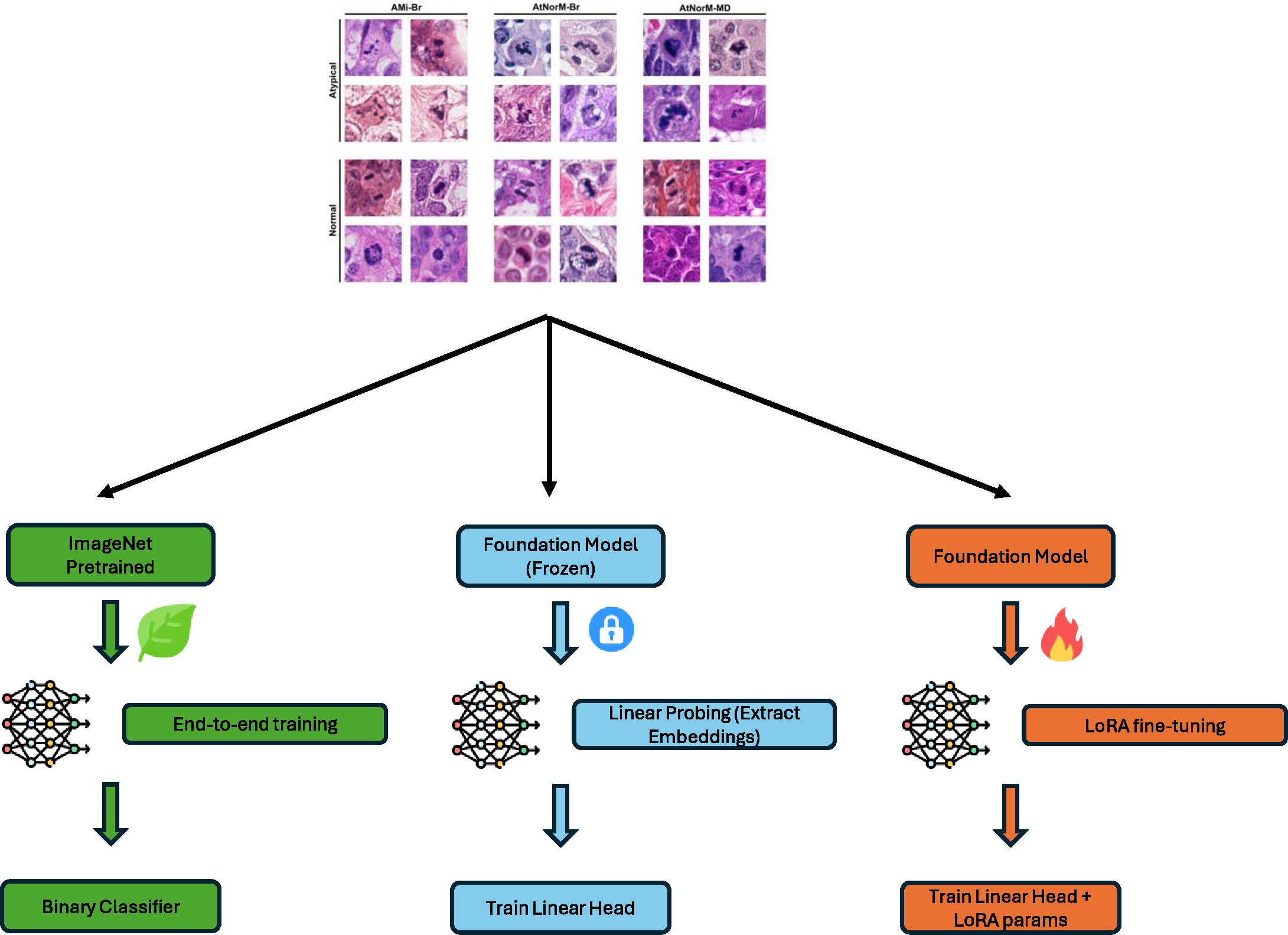
Atypical mitosis marks a deviation in the cell division process that has been shown be an independent prognostic marker for tumor malignancy. However, atypical mitosis classification remains challenging due to low prevalence, at times subtle morphological differences from normal mitotic figures, low inter-rater agreement among pathologists, and class imbalance in datasets. Building on the Atypical Mitosis dataset for Breast Cancer (AMi-Br), this study presents a comprehensive benchmark comparing deep learning approaches for automated atypical mitotic figure (AMF) classification, including end-to-end fine-tuned deep learning models, foundation models with linear probing, and foundation models fine-tuned with low-rank adaptation (LoRA). For rigorous evaluation, we further introduce two new held-out AMF datasets - AtNorM-Br, a dataset of mitotic figures from the TCGA breast cancer cohort, and AtNorM-MD, a multi-domain dataset of mitotic figures from a subset of the MIDOG++ training set. We found average balanced accuracy values of up to 0.8135, 0.7788, and 0.7723 on the in-domain AMi-Br and the out-of-domain AtNorm-Br and AtNorM-MD datasets, respectively. Our work shows that atypical mitotic figure classification, while being a challenging problem, can be effectively addressed through the use of recent advances in transfer learning and model fine-tuning techniques. We make all code and data used in this paper available in this github repository: https://github.com/DeepMicroscopy/AMi-Br_Benchmark
Keywords
Atypical Mitosis · Deep Learning · Foundation Models · Classification · Benchmarking · Histopathology · low-rank adaptation
Bibtex
@article{melba:2026:006:banerjee,
title = "Benchmarking Deep Learning and Vision Foundation Models for Atypical vs. Normal Mitosis Classification with Cross-Dataset Evaluation",
author = "Banerjee, Sweta and Weiss, Viktoria and Donovan, Taryn A. and Fick, Rutger H.J. and Conrad, Thomas and Ammeling, Jonas and Porsche, Nils and Klopfleisch, Robert and Kaltenecker, Christopher C. and Breininger, Katharina and Aubreville, Marc and Bertram, Christof A.",
journal = "Machine Learning for Biomedical Imaging",
volume = "2026",
issue = "MELBA–BVM 2025 Special Issue",
year = "2026",
pages = "115--125",
issn = "2766-905X",
doi = "https://doi.org/10.59275/j.melba.2026-6c1g",
url = "https://melba-journal.org/2026:006"
}
RIS
TY - JOUR
AU - Banerjee, Sweta
AU - Weiss, Viktoria
AU - Donovan, Taryn A.
AU - Fick, Rutger H.J.
AU - Conrad, Thomas
AU - Ammeling, Jonas
AU - Porsche, Nils
AU - Klopfleisch, Robert
AU - Kaltenecker, Christopher C.
AU - Breininger, Katharina
AU - Aubreville, Marc
AU - Bertram, Christof A.
PY - 2026
TI - Benchmarking Deep Learning and Vision Foundation Models for Atypical vs. Normal Mitosis Classification with Cross-Dataset Evaluation
T2 - Machine Learning for Biomedical Imaging
VL - 2026
IS - MELBA–BVM 2025 Special Issue
SP - 115
EP - 125
SN - 2766-905X
DO - https://doi.org/10.59275/j.melba.2026-6c1g
UR - https://melba-journal.org/2026:006
ER -