Quantifying the Efficacy of Deep Learning-Driven Deformable Registra- tion in Multiplexed-Immunofluorescence Imaging for Nucleus Subtype Classification
Gaurav Rudravaram1, Shunxing Bao1, Lucas W. Remedios2, Aravind R. Krishnan1, Michael E. Kim2, Yihao Liu2, Chenyu Gao1, Rendong Zhang2, Bohan Jiang1, Qi Liu3,4, Ken S Lau3,5,6,7, Joseph T. Roland6, Mary K. Washington8, Lori A. Coburn5,9,10,11, Keith T. Wilson5,9,10,11,8, Yuankai Huo2, Bennett A. Landman1,2,12
1: Department of Electrical and Computer Engineering, Vanderbilt University, Nashville, TN, USA, 2: Department of Computer Science, Vanderbilt University, Nashville, TN, USA, 3: Center for Quantitative Sciences, Vanderbilt University Medical Center, Nashville, TN, USA, 4: Department of Biostatistics, Vanderbilt University Medical Center, Nashville, USA, 5: Vanderbilt Center for Mucosal Inflammation and Cancer, Vanderbilt University Medical Center, Nashville, TN, USA, 6: Epithelial Biology Center, Vanderbilt University Medical Center, Nashville, TN, USA, 7: Department of Cell and Developmental Biology, Vanderbilt University School of Medicine, Nashville, TN, USA, 8: Department of Pathology, Microbiology, and Immunology, Vanderbilt University Medical Center, Nashville, TN, USA, 9: Division of Gastroenterology, Hepatology, and Nutrition, Department of Medicine, Vanderbilt University Medical Center, Nashville, TN, USA, 10: Program in Cancer Biology, Vanderbilt University School of Medicine, Nashville, TN, USA, 11: Veterans Affairs Tennessee Valley Healthcare System, Nashville, TN, USA, 12: Department of Biomedical Engineering, Vanderbilt University, Nashville, TN, USA
Publication date: 2026/04/05
https://doi.org/10.59275/j.melba.2026-912a
Abstract
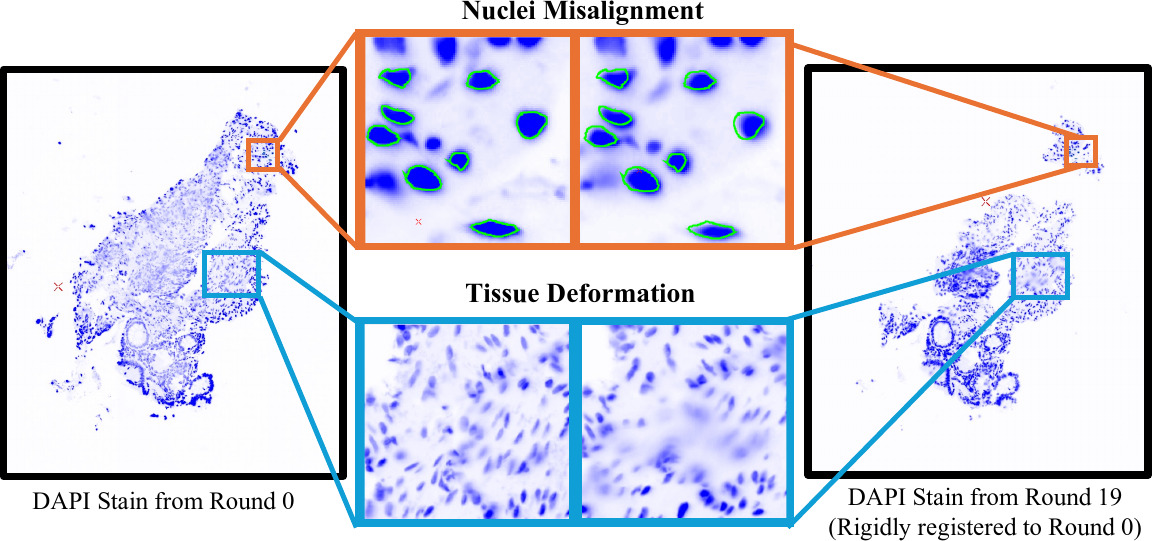
Accurate identification and classification of nucleus subtypes is crucial for cell tracking and uncovering patterns across cell types, such as local cell neighborhoods. Multiplexed immunofluorescence (MxIF) imaging is a process that involves staining, imaging, and then bleaching the same tissue multiple times. Repeating MxIF staining with different marker combinations enables subclassification of cells. However, repeated cycles of staining and bleaching can cause deformation, movement, and tissue loss, resulting in misalignment of markers at the nucleus level. This misalignment can lead to the exclusion of a significant number of cells during downstream analysis. We propose that applying a post hoc deep learning-based deformable registration technique (VoxelMorph) on the respective 4′ ,6-diamidino-2- phenylindole (DAPI) image for each round of staining can reduce the number of nuclei that are excluded due to spatial misalignment across successive staining rounds. By applying the registration transformations from different DAPI rounds to their corresponding stains, we achieve stain registration at pixel-level. To tackle the challenge of large image sizes, we propose a patch-based training and inference strategy. By analyzing residual displacement from bidirectional registrations, we are able to mask out areas in the tissue with high residual displacement to indicate image regions that should not be included for downstream analyses. For validation, we used a deterministic decision tree, based on biological domain knowledge, to classify MxIF nuclei into either one of 13 different classes or an undefined class. Our proposed registration approach effectively reduced the number of undefined nuclei, and we observed a 17.6% increase in the number of successfully classified nuclei compared to a baseline rigid registration. Our code is available at https://github.com/MASILab/MxIF_Registration
Keywords
Machine Learning · Image Registration · Cell Type Classification
Bibtex
@article{melba:2026:010:rudravaram,
title = "Quantifying the Efficacy of Deep Learning-Driven Deformable Registra- tion in Multiplexed-Immunofluorescence Imaging for Nucleus Subtype Classification",
author = "Rudravaram, Gaurav and Bao, Shunxing and Remedios, Lucas W. and Krishnan, Aravind R. and Kim, Michael E. and Liu, Yihao and Gao, Chenyu and Zhang, Rendong and Jiang, Bohan and Liu, Qi and Lau, Ken S and Roland, Joseph T. and Washington, Mary K. and Coburn, Lori A. and Wilson, Keith T. and Huo, Yuankai and Landman, Bennett A.",
journal = "Machine Learning for Biomedical Imaging",
volume = "2026",
issue = "April 2026 issue",
year = "2026",
pages = "186--199",
issn = "2766-905X",
doi = "https://doi.org/10.59275/j.melba.2026-912a",
url = "https://melba-journal.org/2026:010"
}
RIS
TY - JOUR
AU - Rudravaram, Gaurav
AU - Bao, Shunxing
AU - Remedios, Lucas W.
AU - Krishnan, Aravind R.
AU - Kim, Michael E.
AU - Liu, Yihao
AU - Gao, Chenyu
AU - Zhang, Rendong
AU - Jiang, Bohan
AU - Liu, Qi
AU - Lau, Ken S
AU - Roland, Joseph T.
AU - Washington, Mary K.
AU - Coburn, Lori A.
AU - Wilson, Keith T.
AU - Huo, Yuankai
AU - Landman, Bennett A.
PY - 2026
TI - Quantifying the Efficacy of Deep Learning-Driven Deformable Registra- tion in Multiplexed-Immunofluorescence Imaging for Nucleus Subtype Classification
T2 - Machine Learning for Biomedical Imaging
VL - 2026
IS - April 2026 issue
SP - 186
EP - 199
SN - 2766-905X
DO - https://doi.org/10.59275/j.melba.2026-912a
UR - https://melba-journal.org/2026:010
ER -